Townsley helps find COVID-19 vaccine solution one genome sequence at a time
Lipscomb alumni are leading the way to develop tools to aid researchers in finding a vaccine to fight COVID-19.
Kim Chaudoin |

Thomas Townsley (’19) was among a team of bioinformatics researchers who recently released Pantograph to aid researchers in developing a COVID-19 vaccine.
2022 Update: The College of Computing and Technology is now the School of Computing and the School of Data Analytics and Technology.
While the world is feverishly working to develop a COVID-19 vaccination, one Lipscomb alumnus has been working with a group of scientists and researchers from around the world since March to build a pangenomic tool to aid vaccine research.
Last week, Thomas Townsley, who graduated from Lipscomb in December 2019 with a Master of Science degree in software engineering, was among a team of bioinformatics researchers who released PantographTM, a visual browser for Graph Genomes, a new way of capturing sequence data which aids researchers in studying the sequence diversity of the virus which helps in successfully developing a vaccine. The software is free and open source. Townsley says his research team, led by Josiah Seaman, a Ph.D. candidate at Queen Mary University of London, hopes to “get it into the hands of as many researchers as possible” to expedite the development of a COVID-19 vaccination.
The problem that Townsley and his team of researchers have been hard at work to solve is to provide those developing vaccines the most amount of true genetic diversity to generate a selection sweep against COVID-19 when introducing a new vaccine.
“When we are talking about genomes, historically speaking we have worked with a type of data structure called a multiple sequence alignment,” explains Townsley. “What you do in this process is take a lot of different sequences — like my DNA sequence, your DNA sequence and if we sequenced 60 other people’s DNA — and stack all of these DNA sequences on top of each other and then compare them and see how similar or different we are. The one individual we align the 60 sequences to is the linear reference genome which usually isn't a single person but more of a consensus.”
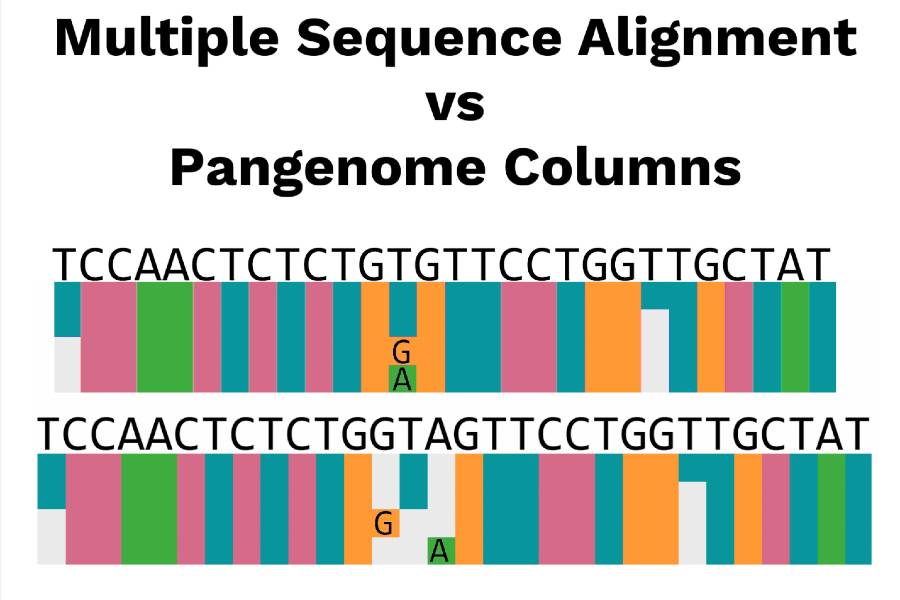

Eliminating Reference Bias in Genomes
Townsley says that what happens is reference bias occurs with respect to aligned individuals and their reference genome. When genomes find a “consensus” the data does not capture the genomic diversity of a population. As a result, reference bias occurs. Because of this, sample reads that highly differ from the reference do not map correctly, and are either mapped to the wrong position in the genome or remain unmapped altogether. Incorrect read mapping in turn leads to false negative or false positive variant calls. This causes problems for researchers as they work to develop solutions or vaccines to serve a greater population.
What Townsley’s group has focused on is identifying real structural variations in SARS-CoV-2. Eliminating reference bias helps us more easily identify these variations by eliminating the false positives. A pangenome is the entire gene set of all strains of a species. It includes genes present in all strains (also known as core genome) and genes present only in some strains of a species, or variable genomes. The core genome represents the genes present in all strains of a species.
“Using our previous illustration, we don’t want to lose information from the 60 individuals’ genomes we sequenced. We want to see how each individual in this group's genes vary from each other but we aren’t losing that information because of reference bias,” says Townsely. “Pangenomes are powerful entities that are going to play a part, particularly for the next 20 years or so, in genetics research especially as we develop drugs that are more targeted than they are now because we have a more precise description of what we are targeting.”
“We have already seen how coronavirus has mutated over time. So the more genetic information we have about coronavirus, the better our vaccines get." — Thomas Townsley (’19)
In the case of coronavirus, Townsley says, proteins in SARS-CoV-2, or what has become known around the world as COVID-19, are highly mutative. He said there are clinical trials for vaccines targeting the Spike protein on the outside of the virus that rely on a lack of genetic diversity. But if there are strains of the mutant protein, the vaccine would be rendered ineffective and the virus would continue to spread. SARS-CoV-2 is an RNA virus that has a much higher mutation capability because of the high number of people it has and continues to infect.
“We have already seen how coronavirus has mutated over time. So the more genetic information we have about coronavirus, the better our vaccines get,” he says. “But we also don’t want to make vaccines that have implicit reference bias in them. So that’s the gist of the problem we are trying to solve.”
Unlocking the Next Level of Population Genetics
The Pantograph project, which began in 2018 before COVID-19 emerged as a way to unlock the next level of population genetics for researchers, quickly became more relevant to the current pandemic. According to the Pantograph website, “The success or failure of our efforts to fight the COVID-19 disease rely upon the sequence diversity of the virus itself. Tests for infection rely on knowing the exact sequence being tested. A rearrangement in the order of genes, even if the content is the same, will return a false negative test.”
The project cites the common cold, also classified as a coronavirus, as an example of a virus that is difficult to eradicate because of the number of people infected and thus the high number of mutations that exist around the world. “Current sequencing techniques may be under-representing the full sequence diversity of the virus because they are reference based. Eliminating reference bias and enabling species genetic diversity on thousands of individuals is the core goal of using a graph genome,” according to the project’s findings.
Townsley said Pantograph is a small piece of the worldwide effort to find a vaccine and to eradicate COVID-19 that can also have a wide-range of disease applications in the future. The short-term goal of the project is to extend the Pantograph tool to add features to make it easier to study the SARS-CoV-2 mutants as the situation develops.
Pantograph has developed the first graph genome browser design with the capability to scale to thousands of individuals and still show the individual’s nucleotide sequence. This means it’s uniquely suited to providing a global overview of species genetic diversity with the option to zoom in on small features.
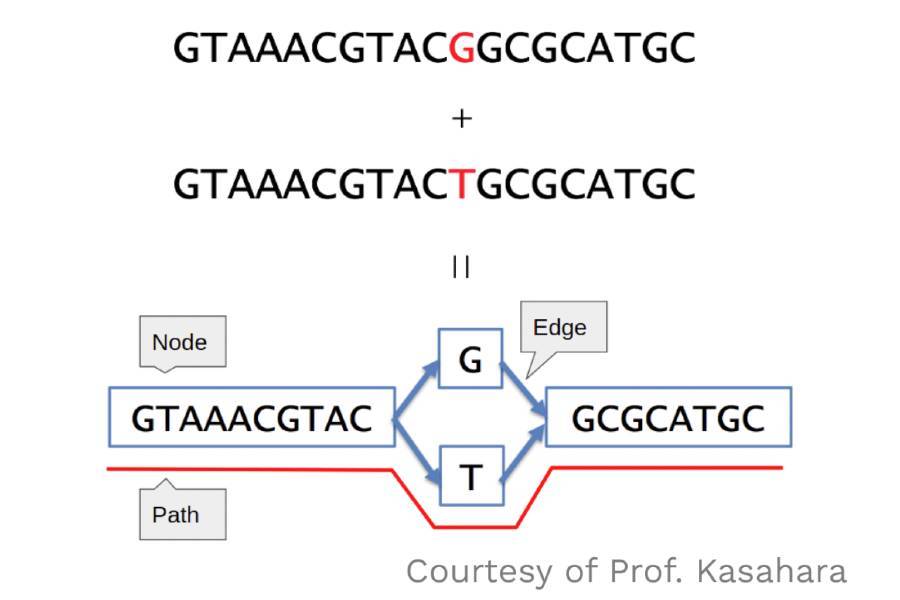
Genome Graph Solution
Townsley explains that linear reference genome representations in current use cannot capture non-reference sequence variation when visualizing hundreds of individuals. Genome graphs are an attractive solution to visualize and investigate the whole variation within large sets of individuals at glance. Structural variants are associated with many diseases, including childhood cancers and schizophrenia. In order to make sequencing data actionable for clinicians, structural variants need to be put in the larger context of all known genetic variation. Understanding genetic variation in plants is equally important for identifying pest and pathogen resistance and increasing crop yields to support a growing population.
The Pantograph project developed Pangenome Schematics, the first visualization with the scalability to render graph genomes from thousands of individuals over gigabase genome sizes with the ability to show both whole chromosome features as well as zoom into nucleotide sequence variation. Townsley says that no tool to date has satisfied all these criteria. Scalability is accomplished through graph sorting and binning adjacent sequences to create shared syntenic blocks called components. Non-linear structural variants can be shared by many individuals as links across the pangenome.
Townsley joined the core project team in March as the point person responsible for maintaining the graph genome pipeline and as the release engineer to push the product out to researchers.
“Our aim is to get this in the hands of comparative genome researchers, and that extends to vaccine research,” he says. “Our hope is that anyone who has an interest in really studying the genetic variations of coronavirus can use our software to pick up on variations in coronavirus that they would not be able to see otherwise due to the limitation of other multiple sequence alignment technologies.”
The PantographTM pangenome graph browser for SARS-CoV-2 may be viewed at graphgenome.org.
“Our aim is to get this in the hands of comparative genome researchers, and that extends to vaccine research. Our hope is that anyone who has an interest in really studying the genetic variations of coronavirus can use our software to pick up on variations in coronavirus that they would not be able to see otherwise due to the limitation of other multiple sequence alignment technologies.” — Thomas Townsley
Making a Difference in the World
Being part of a project that has the potential to have a positive impact on the lives of people around the world is very satisfying for Townsley.
A Sulphur, Louisiana native who earned his undergraduate degree from Loyola University in 2016, Townsley’s background is in management and information systems. He says science has always been interesting to him. After getting a “good job” out of college in the corporate world, Townsley says he didn’t feel like he was making a contribution to society. He decided to return to school and saw that Lipscomb University was one of the few institutions that had an interdisciplinary program around computational biology.
“So I reached out to Lipscomb’s College of Computing & Technology,” he recalls. “They were super nice and I told them my story. I was fortunate to work for a couple of years in a good job. But I quit my job and basically dove right into my studies at Lipscomb.”

Joe Deweese, Associate Professor, Pharmaceutical Sciences
In 2019, prior to graduation, Townsley was a supported research assistant in the College of Computing & Technology with Tim Wallace, associate professor and chair of the Department of Computational Sciences, engaged in a new research collaboration effort lead by Joe Deweese, associate professor in the Department of Pharmaceutical Sciences in the College of Pharmacy.
After graduating from Lipscomb in December 2019, Townsley had the opportunity to remain at Lipscomb to continue on as the team’s software developer. For the past year Townsley has worked as part of Deweese’s research group working on a software program that will predict features of protein structure using the amino acid sequence of a protein. This tool will be helpful in the development of new drugs and in studying new proteins.
"Thomas has worked with my research collaborators and I over the last year in developing a new software tool to study protein structure,” says Deweese. “Thomas is a very talented programmer, and he continually comes up with creative ways to solve our programming challenges. We are currently testing our software tool before we prepare to release it for use by researchers worldwide."
Deweese is friends with Josiah Seaman, who is overseeing the Pantograph project. “When Josiah put out the call for programmers, I knew Thomas would be the right guy for the job,” recalls Deweese.
It was a good fit for Townsley who had been interested in “doing something in the COVID space.” Townsley has returned to the College of Pharmacy research project now that he has completed his commitment to the Pantograph project.
“I’m more of a computer person but I definitely enjoy working with biologists and scientists because they are so passionate about what they do,” he admits. “Everything they work on has an immediate quality to it - like this could actually help us target cancer-causing entities. That is really motivating to me coming from a corporate background. I’d much rather be working on something like that where I feel like I’m making a meaningful difference.”
Having been part of two major research projects in recent months, Townsley discovered first-hand that “science can be really exciting and you don’t necessarily need to be a biologist in a lab to make a meaningful contribution,” he says.
“I don’t consider myself a biologist and I never thought in a million years that I would be working on a problem like this. I had this idea before that science was a niche thing and that only a select few people would actually be able to be a part of it. Everyone actually does have a role that they can play.”
To learn more about the Pantograph project, visit https://graph-genome.github.io/.
Learn more about Lipscomb’s College of Computing & Technology at www.lipscomb.edu/computing.